Quantitative mass spectrometry as a powerful tool for analysing chitosan-modifying enzymes and how they define the bioactivities of chitosans
Stefan Cord-Landwehr
Supervisor: Prof. Dr. Bruno Moerschbacher
Co-Supervisor: Prof. Dr. Henk Schols
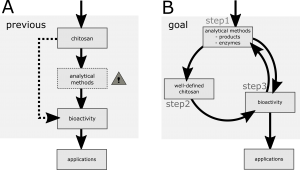
Chitosans are a group of highly promising biomolecules, which consist of β(1-4) linked glucosamine (GlcN; D) and N-acetylglucosamine (GlcNAc; A) units. They have a wide variety of biological activities and therefore many potential applications in e.g. agriculture or medicine. It is known that their biological activities depend on the degree of polymerization (DP) and acetylation (DA). Additionally the pattern of acetylation (PA) likely influences these properties. However, the lack of well-defined partially acetylated chitosan oligomers (paCOS) and the time- and material-consuming and sometimes poorly used analytical methods are some of the main obstacles to understanding their structure-function relationship and therefore the application of chitosans (Scheme A).
The studies presented in this doctoral thesis have the goal of improving each part of the process required for understanding and using chitosans, which will thereby reinforce the connections between these parts, allowing us to gain more information about each aspect (Scheme B). To achieve these goals, the first step (Scheme B) is to improve and develop methods to analyse the chitosan products themselves; through this, we can learn more about the chitin- and chitosan-modifying enzymes (CCME) used to produce them. The second step is to produce novel and well-defined chitosans, and then perform in the third step bioactivity tests on these compounds. In turn, this offers information about the natural functions of the initial enzymes used to produce the paCOS products (return to step 1, Scheme B), as well as potential applications for chitosan.
To address the first step (Scheme B), a hydrophilic interaction liquid chromatography-mass spectrometric (HILIC-MS) method developed to analyse plant cell wall carbohydrates was optimised to analyse the fully acetylated chitin, the fully deacetylated and partially acetylated chitosan oligosaccharides within one experiment. Quantitative analysis of paCOS including their DP, DA, and PA was achieved by (I) [2H3]N-acetylation of free amino groups in the paCOS, (II) spiking the samples with [13C2; 2H3]acetylated chitin oligomers as internal standards, (III) analysing the mixture by HILIC-ESI-MS, and (IV) pattern determination using HILIC-ESI-MS/MS with paCOS containing an 18O label at the reducing end. The main outcome of this quantitative HILIC-ESI-MSn analysis is the fast, efficient and precise determining of subsite and substrate specificity of chitin- and chitosan-modifying enzymes (CCME) while only requiring a small amount of produced oligosaccharides compared to the previous used SEC-NMR method. Using ChiB from Serratia marcescens, the best-described chitinase, as benchmark, the newly developed quantitative HILIC-ESI-MS method was verified as having only small variations compared to the results obtained with the common SEC-NMR method. Furthermore, the increased sensitivity of the method enabled the monitoring of the subsite specificity at different time points and for chitosans with different DAs. Additionally the new quantitative analysis HILIC-ESI-MS method allows for investigation of the interplay between the neighbouring subsites (-2 and -1, +1 and +2); in contrast, the SEC-NMR method only allows one subsite isolated from the neighbouring units to be studied.
Subsequently, studies which addressed step 2 in Scheme B, were performed with a novel chitin deacetylase (CDA) derived from the endophytic fungus Pestalotiopsis sp. and its products. This enzyme (PesCDA), recombinant produced in E. coli as MBP fusion protein, is a regio-selective CDA and generates novel and well-defined paCOS with the pattern
AA[D]n-3A. To address step 3 (Scheme B) bioactivity analyses on the products of PesCDA were performed, showing that this enzyme converts by deacetylation the elicitor-active chitin oligomer into an elicitor-inactive molecule. This inactivation of chitin oligomers supports a long-standing hypothesis for how chitin-containing fungi overcome detection by the plant cells and therefore avoid evoking the plant’s immune system response. As such, this might be the natural function of PesCDA for the endophytic fungus, revealed by performing bioactivity tests with the products of this enzyme. Furthermore, a subsequently detailed analysis of PesCDA subsite specificity allowed knowledge-based combination of PesCDA with other regio-selective CDAs to produce additional well-defined paCOS with new pattern of acetylation, which can be used for bioactivity tests in the future.
The studies shown in this thesis provide several new possibilities to analyse chitin and chitosan oligosaccharides with a focus on quantification to enable a precise characterisation of chitin and chitosan-modifying enzymes and their chitosan products. Based on this, various chitin- and chitosan-modifying enzymes can be characterised, and several well-defined chitosans can be produced which can be used now for bioactivity assays to decipher the structure-function relationships of chitosans. The future studies should not only be oxidative burst studies, as shown for the PesCDA products with rice cell suspension cultures, but should be more significant for the biotechnological usage, such as by testing chitosan’s effects in plant promotion or plant strengthening. By interconnecting each part of the chitosan studies, we will be able to obtain more refined knowledge about chitosans and the enzymes used to produce them, which will therefore enable more and more applications.
Publications
1. Cord-Landwehr, S.; Ihmor, P.; Niehues, A.; Luftmann, H.; Moerschbacher, B. M.; Mormann, M., Quantitative Mass-Spectrometric Sequencing of Chitosan Oligomers Revealing Cleavage Sites of Chitosan Hydrolases. Analytical Chemistry, (2017) 89, 2893-2900; < http://dx.doi.org/10.1021/acs.analchem.6b04183 >.
2. Naqvi, S.; Cord-Landwehr, S.; Singh, R.; Bernard, F.; Kolkenbrock, S.; El Gueddari, N. E.; Moerschbacher, B. M., A Recombinant Fungal Chitin Deacetylase Produces Fully Defined Chitosan Oligomers with Novel Patterns of Acetylation. Applied and Environmental Microbiology, (2016) 82, 6645-6655; < http://dx.doi.org/10.1128/aem.01961-16 >.
3. Cord-Landwehr, S.; Melcher, R. L. J.; Kolkenbrock, S.; Moerschbacher, B. M., A chitin deacetylase from the endophytic fungus Pestalotiopsis sp. efficiently inactivates the elicitor activity of chitin oligomers in rice cells. Scientific Reports, (2016) 6; < http://dx.doi.org/10.1038/srep38018 >.
4. Langner, T.; Oeztuerk, M.; Hartmann, S.; Cord-Landwehr, S.; Moerschbacher, B.; Walton, J. D.; Goehre, V., Chitinases Are Essential for Cell Separation in Ustilago maydis. Eukaryotic Cell, (2015) 14, 846-857; < http://dx.doi.org/10.1128/ec.00022-15 >.
5. Hamer, S. N.; Cord-Landwehr, S.; Biarnes, X.; Planas, A.; Waegeman, H.; Moerschbacher, B. M.; Kolkenbrock, S., Enzymatic production of defined chitosan oligomers with a specific pattern of acetylation using a combination of chitin oligosaccharide deacetylases Scientific Reports, (2015) 5; < http://dx.doi.org/10.1038/srep08716 >.
6. Remoroza, C.; Cord-Landwehr, S.; Leijdekkers, A. G. M.; Moerschbacher, B. M.; Schols, H. A.; Gruppen, H., Combined HILIC-ELSD/ESI-MSn enables the separation, identification and quantification of sugar beet pectin derived oligomers. Carbohydrate Polymers, (2012) 90, 41-48; < http://dx.doi.org/10.1016/j.carbpol.2012.04.058 >.